Sequence
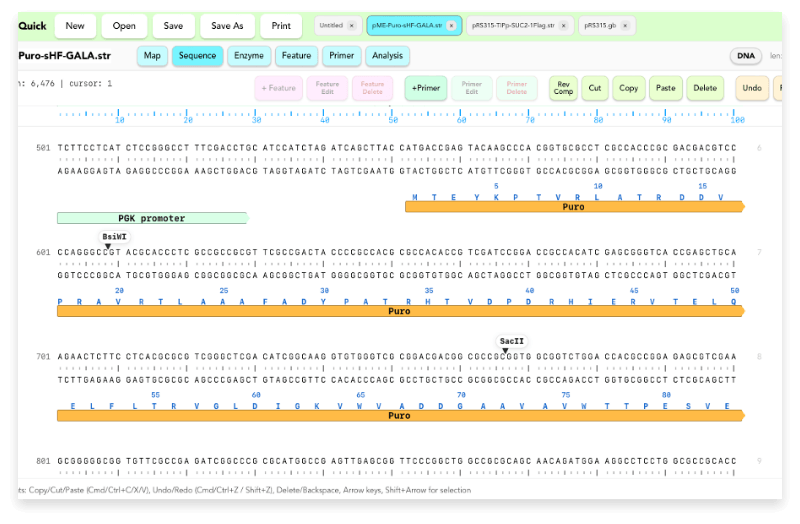
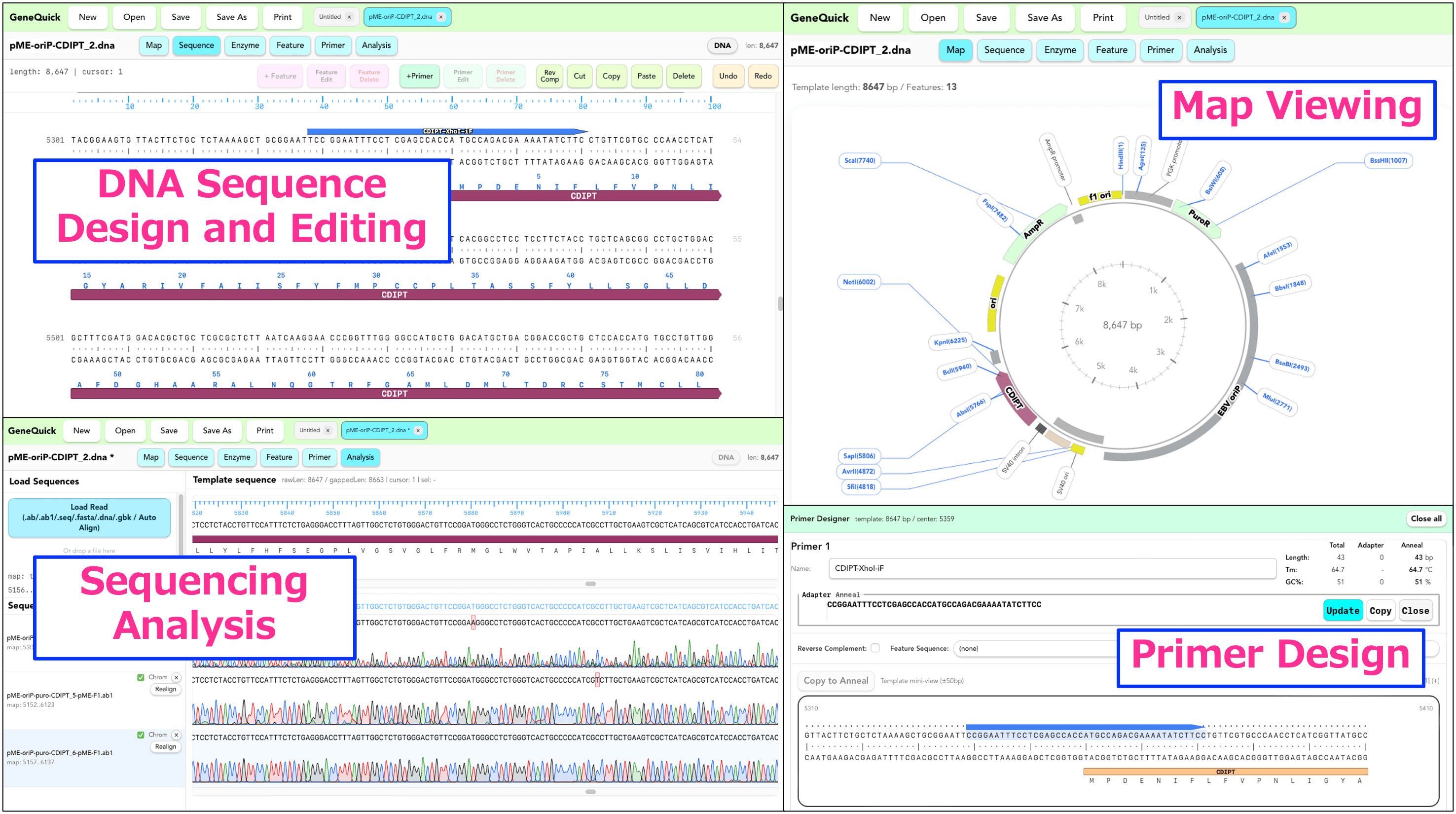
This is the main screen for displaying and editing nucleotide sequences. It supports copy, cut, paste, delete, replace, reverse complement, and Undo / Redo, while also connecting naturally to feature and primer editing.
- Direct nucleotide sequence editing
- DNA / RNA mode switching
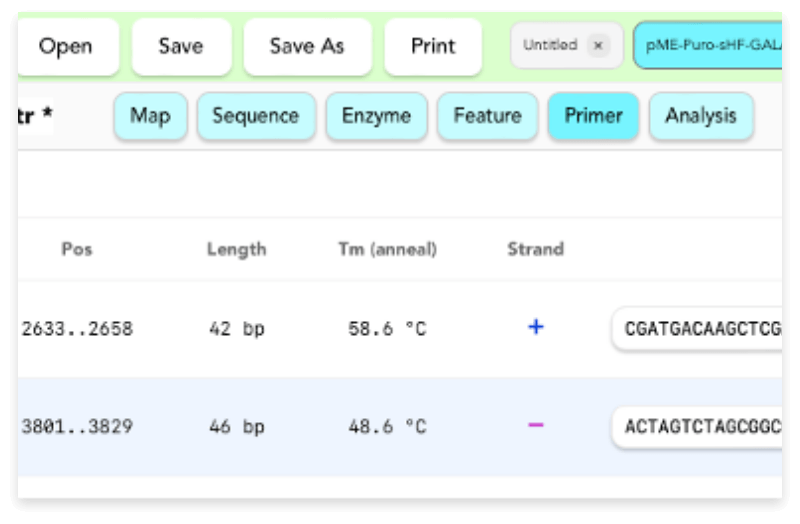
- Selecting and editing Features / Primers
- Amino acid sequence copy support